Count matrix analysis - application to molecular microbial data
Methods
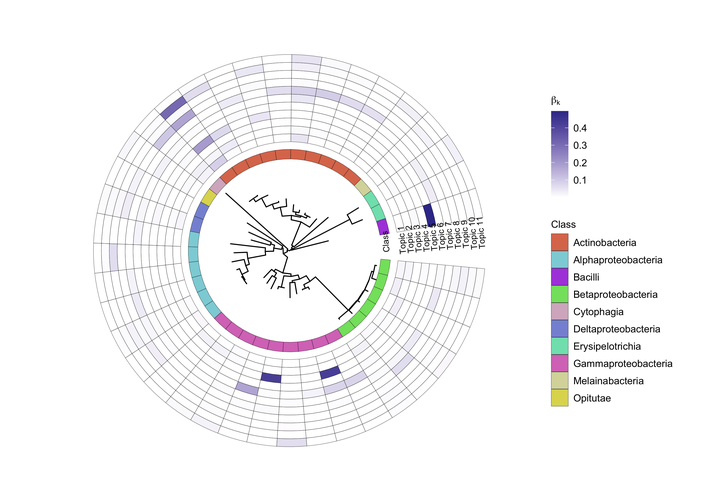
Differential topic analysis
In microbiome research, an important goal is often to find taxonomic differences across environments or groups. We introduce differential topic analysis that facilitates inferences on latent microbial communities.
- Software
- Papers and Preprint
- A Statistical Perspective on the Challenges in Molecular Microbial Biology, Journal of Agricultural, Biological and Environmental Statistics.
- Talks
- BioC 2021, August 4, 2021. Slides.
- R-Ladies Dallas, July 12, 2021. Slides.
- International Conference on Environmental and Medical Statistics, January 9, 2020.
DNA contamination removal in molecular microbial studies
Molecular technologies can quantify bacteria in low biomass samples, such as blood. However, DNA contamination from external sources misidentifies the taxon’s provenance. We developed a Bayesian reference analysis to infer DNA contamination.
- Software
- Papers and Preprint
- Method paper (To be submitted).
- Talks
- Statistics Seminar, Department of Statistics, Stanford University - June 2, 2020.
- 21st Meeting of New Researchers in Statistics and Probability - July 24-27, 2019.
- 3rd Workshop on Statistical and Algorithmic Challenges in Microbiome Data Analysis - April 2, 2019.
Inference on longitudinal microbiome data
Longitudinal designs help experimenters overcome some of the difficulties caused by the temporal and subject-to-subject variability of the microbiome. They also allow subjects to be used as their own controls.
The proposed resampling method combined moving block bootstrap (MBB) method, empirical subsampling method, mixture model, generalized linear model, generalized estimating equation, median-ratio method, and shrinkage estimation to enabling inference on microbiome longitudinal data. With the optimal block size computed using subsampling, the MBB method accounts for within-subject dependency by using overlapping blocks of repeated observations within each subject to draw valid inferences based on approximately pivotal statistic.
- Software
- Papers and Preprint
- Talks
- Biomedical Computation at Stanford 2018, April 19, 2018.
Applications
diffTop
- In progress…
BARBI
- Application to identify translocation of bacteria (To be submitted)
- Combined use of metagenomic sequencing and host response profiling for the diagnosis of suspected sepsis. BioRxiv
BootLong
- Identify deferentially abundant taxa in preterm and term labor (vaginal microbiome data).